HKUST Researchers Develop Efficient and Accessible Single-Molecule Platform for Detecting Various Amylin Species Associated with Type 2 Diabetes
A research team led by The Hong Kong University of Science and Technology (HKUST) has developed an optical plasmonic tweezer-controlled Surface-Enhanced Raman Spectroscopy (SERS) platform that utilizes on-and-off control of light to probe various amylin species in mixtures at the single-molecule level, unveiling the hetergogerous structures of pH-dependent amylin species, and the secrets behind amyloid aggregation mechanisms associated with type 2 diabetes.
By removing ensemble averaging, single-molecule techniques discern the signal of individual molecules to unveil hidden details and revolutionize our understandings of complex and heterogeneous molecular systems. Current single-molecule approaches are limited to ultra-dilution and/or molecular immobilization because the diffraction-limited detection volume cannot be further reduced. Whereas, certain biomolecules engage in various interactions that are significantly affected by concentrations. For example, as a typical intrinsically disordered protein, human Islet Amyloid Polypeptide (amylin, hIAPP) lacks stable secondary structures but possesses aggregation propensity governed by environmental factors, such as concentration and pH, to form different oligomeric intermediates and amyloid fibrils in type II diabetes patients. The molecular mechanism remains unclear, due to the challenges of detecting the rare, transient, and heterogeneous amylin species in a dynamic mixture, calling for the development of advanced single-molecule methods.
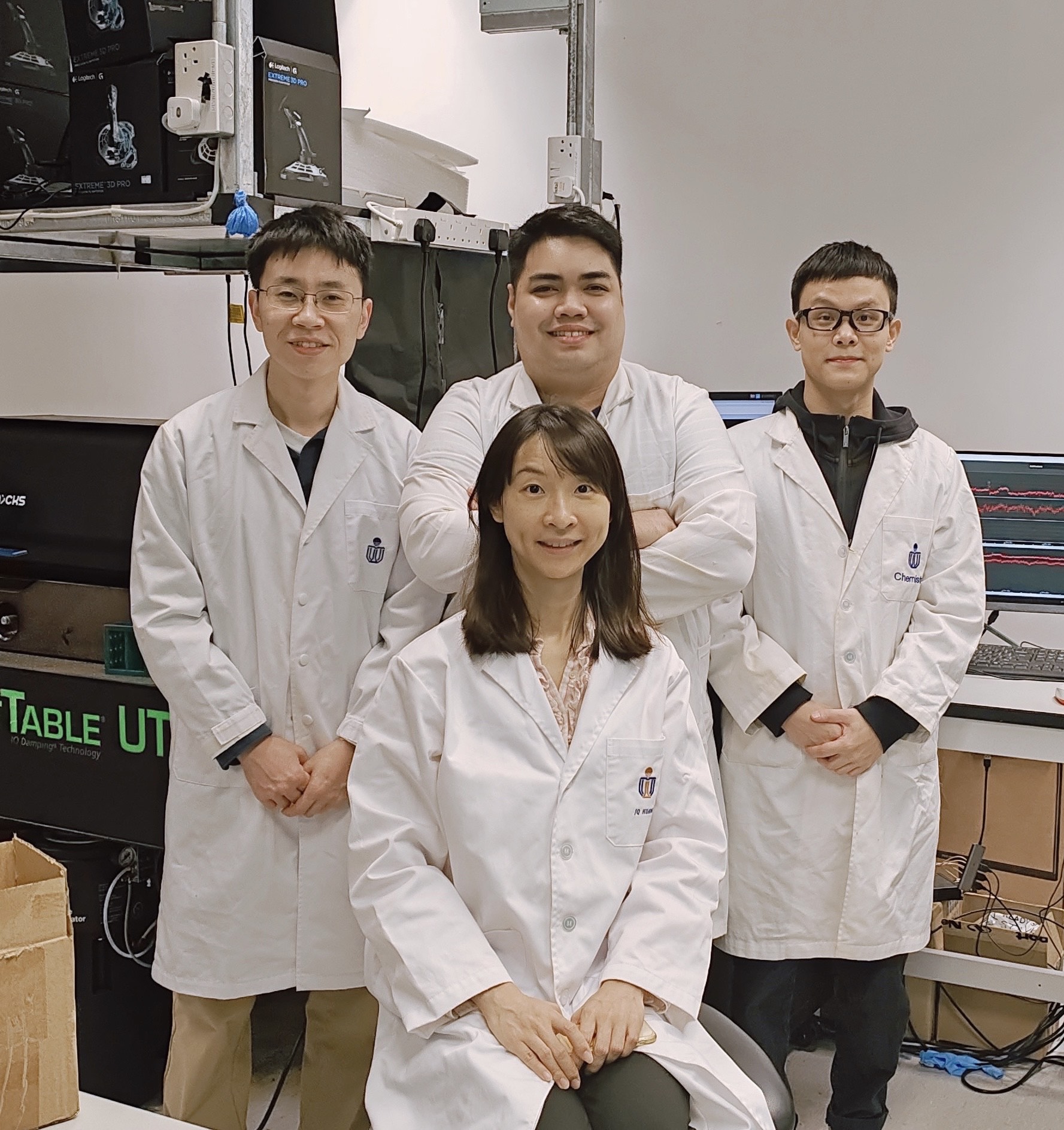
In a recent breakthrough, the research team led by Prof. HUANG Jinqing, Assistant Professor at HKUST’s Department of Chemistry has successfully developed a novel single-molecule platform combining optical plasmonic manipulation and SERS measurement to reduce detection volume and elevate signal enhacement, enabling efficient and high-throughput single-molecule characterization to study pH-dependent amylin species at physiological concentrations.
Specifically, the team constructed a plasmonic junction between two Ag nanoparticle-coated silica microbeads to trap an additional Ag nanoparticle to form a dynamic nanocavity upon laser irradiation, which could encapsulate a single or a few molecules for sensitive SERS characterizations. Since both optical plasmonic trapping and SERS phenomena are spatially confined in the nanometer-scale, it surpasses optical diffraction limit to enable precise position control, minimize detection volume, and boost SERS enhancement simultaneously. Besides, the constructed Ag nanoparticle-coated silica microbead dimers are more stable than the conventional Ag nanoparticle assembles in solutions, making it easier to observe and locate the plasmonic junction on regular microscopes to improve efficiency and reproducibility. By switching the laser light between “on” and “off” states, the researchers can control the optical plasmonic trapping to modulate the assembly and disassembly of the dynamic nanocavity for high-throughput sampling and simultaneous SERS measurements.
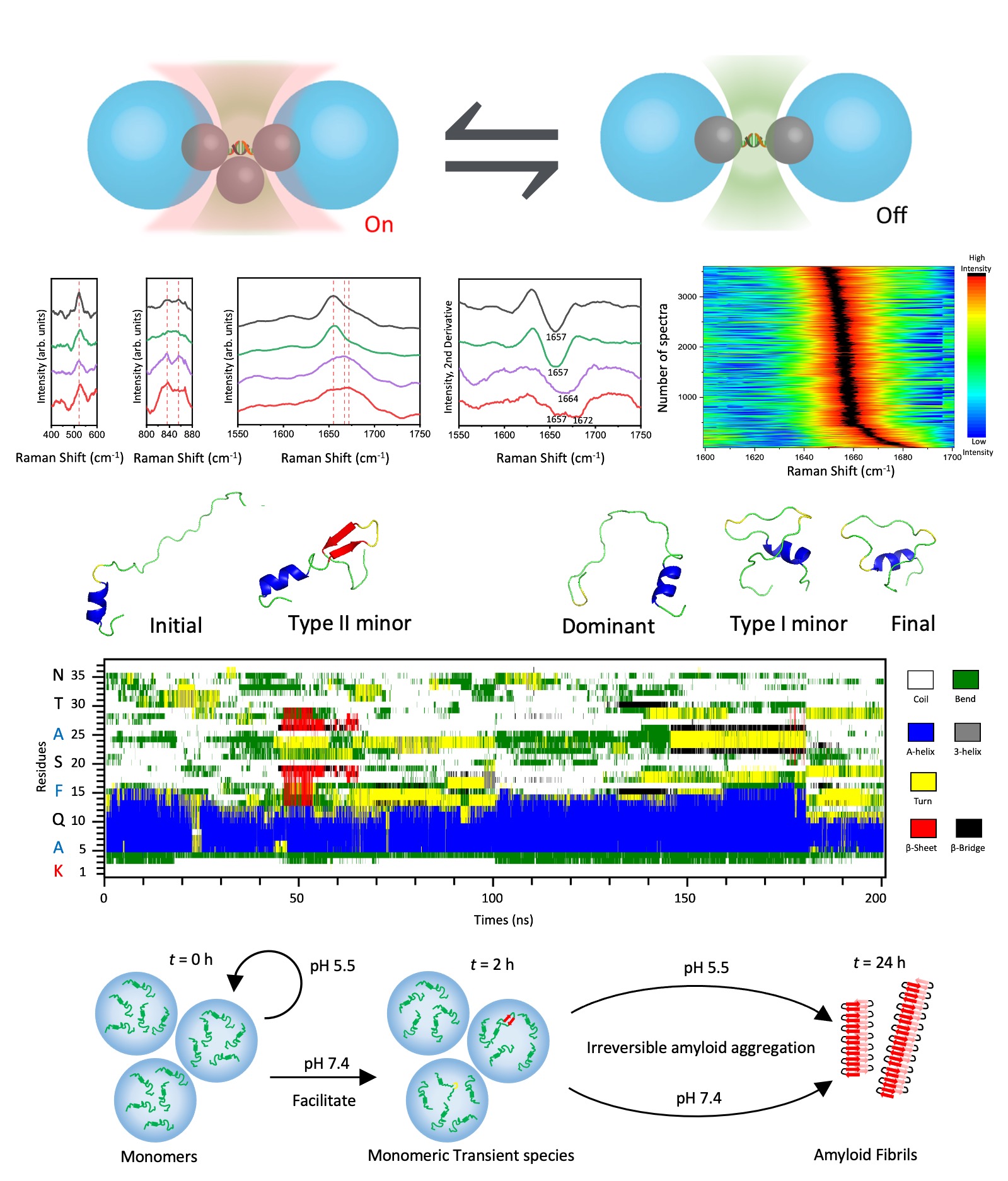
Using this efficient single-molecule platform, the research team harnessed a statistically significant amount of SERS spectra underlying the structural features of various amylin species under two distinct physiological conditions, the secretory granules of pancreatic β-cells at pH 5.5 and the extracellular compartments at pH 7.4, respectively. Two types of low-populated amylin species were identified from its predominant monomers at the early stage of amyloid aggregation in neutral pH, containing a critical turn structure or a short β-hairpin with constrained C-terminal as supported by molecular dynamics (MD) simulations. Such a slight shift in the equilibrium among different amylin species could drive irreversible amyloid development even after the post-adjustment of pH from 7.4 to 5.5. Hence, the direct structural characterization of these amylin species within heterogeneous mixtures indicates the impact of pH on their intra- and inter-molecular interactions and sheds light on the mechanism behind pH-regulated amyloid aggregation for understanding type 2 diabetes.
“We present an easy-to-use strategy that reduces detection volume, enhances molecular signal, and increases turnover efficiency,” explained Prof. Huang. “Our single-molecule platform can acquire a large amount of SERS spectra as molecular snapshots, comparable to those obtained through MD simulations. By statistically analyzing the structural details at the single-molecule level, we are able to reconstruct the bulk properties and gain unique insights into the population and probability of specific molecule types within the heterogeneous mixture. It has the potential to uncover hidden mysteries in complex systems.”
The study was recently published in the scientific journal Nature Communications.